Solid Tumor Profiling
Molecular Profiling of Solid Tumors
The advent of molecular profiling overcame the limitations of traditional solid tumor classification methods, which relied on the morphology of tumor cells and the surrounding tissue.1 Today, molecular profiling is a standard technique for classifying solid tumors, with established guidelines from the College of American Pathologists (CAP)2 and the European Society for Molecular Oncology (ESMO).3
In turn, genomic technology has evolved to meet molecular profiling needs. Next-generation sequencing (NGS) provides a comprehensive method for assessing the majority of genes associated with solid tumors, including lung, colon, breast, melanoma, gastric, and ovarian cancers. NGS also delivers high sensitivity to capture tumor heterogeneity, which other approaches such as Sanger sequencing often miss.4
Benefits of NGS
Tumor profiling using NGS analyzes a select set of genes, gene regions, or amplicons based on known involvement with solid tumors. This targeted approach delivers the high sensitivity to detect rare mutations and tumor subclones, offering increased visibility into important driver mutations in cancer.
Answering Multiple Questions with One Assay
Dr Ravindra Kolhe uses Illumina NGS to study a wide number of variants associated with many different solid tumors.
Rapid NGS Workflow
Solid tumor profiling using NGS follows a simple workflow that can be easily scaled to hundreds of samples. Clinical research labs can go from DNA to report in approximately 40 hours. This workflow uses a single sequencing instrument, minimizing laboratory validation steps for seamless integration into the clinical research lab setting.
Multi-Drug Approaches to Precision Oncology Research
Clinical researchers at Avera Health use DNA and RNA sequencing to analyze solid tumors and uncover multiple drug-susceptible tumor pathways.
Featured Tumor Profiling Articles
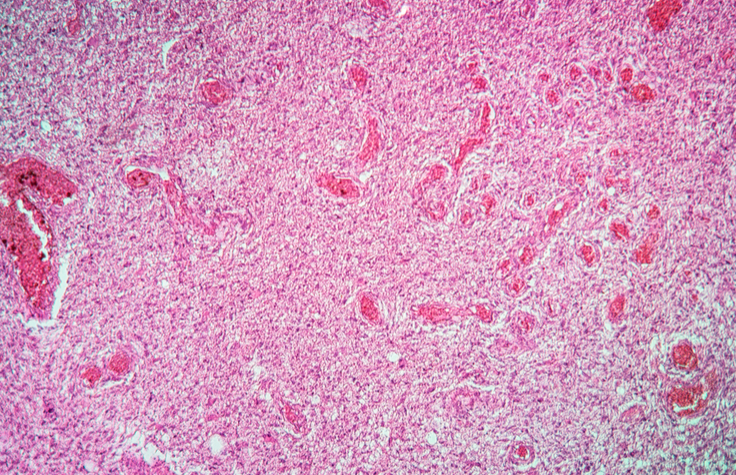
NGS Panels Demonstrate Value in Solid Tumor Studies
In a comparison study, TruSight Tumor 170 accurately identified glioma genetic markers, supporting its use in tumor characterization for clinical research.
Read Customer Interview
Exploring the Genetic Basis of Oral Cancer
Dr. Majumder uses NGS to study tumor heterogeneity, somatic mutations, and germline variants associated with a deadly oral cancer.
Read Customer Interview
Fighting For My Life, I Also Fought For Yours
Sandra Balladares, PhD, talks about her journey to the United States to bring genetic testing to Hispanic breast cancer patients in Latin America.
Read ArticleNGS Workflow Finder
Take the guesswork out of your next workflow. The NGS Workflow Finder provides personalized solution recommendations and resources so you can sequence with confidence.
Find your NGS workflow todayOur Products
Illumina products empower clinical cancer research labs to use the most current technologies to find accurate answers while seamlessly integrating NGS into lab procedures. Library preparation kits are compatible with archival solid tumor tissues and can be easily scaled to process large sample volumes.
Illumina sequencers deliver industry-leading sequencing, generating ~90% of the world’s sequencing data.* Efficient reporting capabilities enable labs to annotate, classify, and communicate significant findings in a standard format.
Click on the below to view products for each workflow step.
Comprehensive panel detects single-nucleotide variants (SNVs), amplifications, and fusions that contribute to solid tumor progression.
TruSight Tumor 15Focused panel to assess relevant solid tumor somatic variants in a simple, sample-to-data workflow.
TruSight RNA Pan-CancerTargeting 1385 cancer-associated genes for gene expression, as well as variant and fusion detection in most oncology sample types including FFPE.
TruSight RNA FusionA gene fusion detection panel targeting fusion associated genes in many cancer types with the ability to detect known and novel fusion gene partners.
TruSight Oncology 500Assay targeting multiple variant types, including tumor mutational burden (TMB) and microsatellite instability (MSI), even from low-quality samples.
Targeted research panel investigating hotspot regions of 50 genes with known associations to cancer.
AmpliSeq for Ilumina Focus PanelTargeted DNA and RNA research panel investigating 52 genes with known relevance to solid tumors.
AmpliSeq for Illumina Comprehensive Panel v3Targeted DNA and RNA research panel investigating variants across 161 genes associated with a range of cancer types.
AmpliSeq for Illumina Comprehensive Cancer PanelTargeted research panel investigating the exonic regions of 409 genes with known associations to cancer.
NGS Target EnrichmentTarget regions of the genome that are relevant to specific research interests.
Simplest and most affordable solution for low-throughput targeted sequencing of solid tumors.
MiSeq SystemDesktop sequencer featuring a simple NGS workflow, integrated data analysis software, and unmatched accuracy.
FDA-cleared in vitro diagnostic NGS system that also operates in research mode to generate accurate, reliable data.
NextSeq 1000 & 2000 SystemsThese groundbreaking sequencers offer flexibility for current and emerging applications, our simplest workflow yet, and data analysis in as little as 2 hours.
Easy-to-use software for analysis and variant calling on the MiSeq System.
Variant AnalysisPowerful analysis and reporting tools that provide biological insight into genomic variant data.
The Illumina genomics computing environment for NGS data analysis and management.
Local Run ManagerEasy-to-use software for automated on-instrument data analysis.
Learn More

Removing Cancer's Veil: TruSight Tumor 170
This enrichment-based cancer research assay interrogates both DNA and RNA, detecting small variants, gene amplifications, gene fusions, and splice variants.

NGS Complements Traditional Cytogenetic Methods
Genomic analysis of tumors provides comprehensive detection of genetic abnormalities.
Interested in receiving newsletters, case studies, and information on cancer genomics?
Sign Up
Sequencing FFPE Samples to Identify Cancer Drivers
Dr. Yasser Riazalhosseini preserves the integrity of FFPE samples to analyze disease-causing mutations and prognostic signatures.
Comprehensive Panel for Gene Fusion Detection
Ravindra Kolhe, MD, PhD, discusses using the TruSight RNA Fusion Panel to study gene fusions in cancer.

Integrative Genomics to Inform Cancer Prognosis
Molecular profiling that combines DNA- and RNA-level data can more easily identify genes that may be driving cancer.
References
- Cross D, Burmester JK. The promise of molecular profiling for cancer identification and treatment. Clin Med Res. 2004;2:147-150.
- Lindeman NI, Cagle PT, Beasley MB, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Thorac Oncol. 2013;8:823-859.
- Mosele F, Remon J, Mateo J, Westphalen CB, et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Ann. Oncol. 2020;31(11):1491-1505.
- Jamuar SS, Lam AT, Kircher M, et al. Somatic mutations in cerebral cortical malformations. N Engl J Med. 2014; 371:733–43.
* Data calculations on file. Illumina, Inc. 2015.
