Rare Disease Genomics
Committed to ending diagnostic odysseys for rare disease
Technology innovation and collaboration in health care to ensure families receive a diagnosis early
Rare disease genomics and precision medicine
Understanding the genomics of rare disease can help doctors pinpoint the cause of undiagnosed disorders, helping families avoid years of hospital visits and unnecessary tests. There are more than 7,000 known rare diseases1 and more discovered every year. Collectively, 2–6% of the population (> 150 million people) is affected by a rare disease.1–3
On average, the long search for a rare disease diagnosis—the “diagnostic odyssey”—takes 5 to 7 years,4 up to 8 physicians,5 and 2 to 3 misdiagnoses.5 Given that 80% of rare diseases are genetic or have a genetic component, comprehensive genomic sequencing increases the potential of uncovering an underlying etiology in patients.6 Next-generation sequencing (NGS) offers the highest likelihood of rare disease diagnosis7–8 and the fastest path to ending the diagnostic odyssey.7
No rare disease will go unseen
Nothing exists—until it is named. Be part of the collective mission to end the diagnostic odyssey through rare disease genomics.
View VideoThe value of genomics for rare disease diagnosis
Genomics is driving a fundamental shift in rare disease diagnosis, from symptom analysis to molecular etiology assessment. Understanding the biological basis of disease can lead to better care and targeted treatment, with predictable, evidence-based outcomes. This type of molecular diagnosis in rare disease genomics is the basis for precision medicine.
Molecular diagnosis of rare disease is a critical step that can benefit patients, their families, physicians, and other care providers. According to the American College of Genetics and Genomics (ACMG), the identification of the genetic etiology of an individual’s disease has utility for the patient, their family, and society at large. 9
The genome as a backbone for molecular analyses
Learn why whole-genome sequencing offers value as a backbone to any molecular genomics assay.
View VideoBenefits of rare disease genomics
Tailored Disease Management
An understanding of rare disease mechanisms allows physicians to refer patients to appropriate specialists, select tailored therapeutics, and offer disease-specific follow-up.
Reduced Expenses
By avoiding lengthy diagnostic odysseys, genomic diagnoses for rare disease can help prevent costly tests and procedures and limit unnecessary referrals.
Reproductive Counseling
Defining the inheritance pattern of a rare disease informs recurrence risks for patients and both their immediate and extended families, supporting informed family planning.
Psychosocial Benefits
In addition to avoiding the stress associated with diagnostic odysseys, receiving a molecular diagnosis brings affected families together in a community of rare disease support groups.
Societal Benefits
Understanding the genomics of rare disease can help identify new drug targets and improve the efficiency of care.
Rare Disease Patient Stories
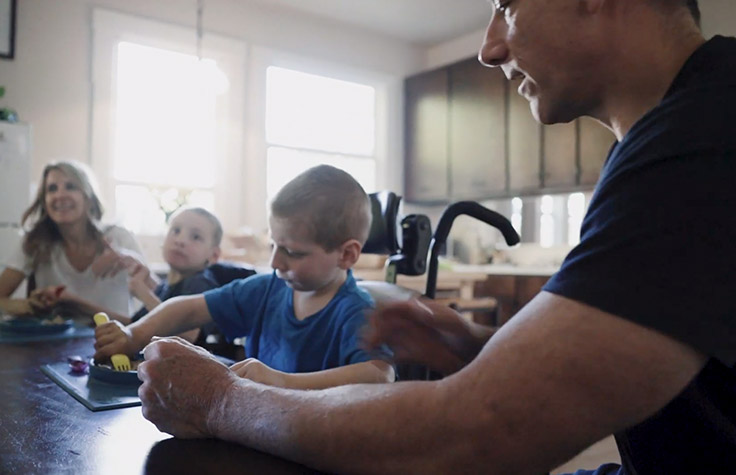

Ending Carson & Chase’s 6-year diagnostic odyssey
Carson was initially diagnosed with cerebral palsy, but his brother, Chase, proved that to be a misdiagnosis. After four more years, whole-genome sequencing helped diagnose Carson and Chase definitively with mitochondrial enoyl CoA reductase protein-associated neurodegeneration (MEPAN).
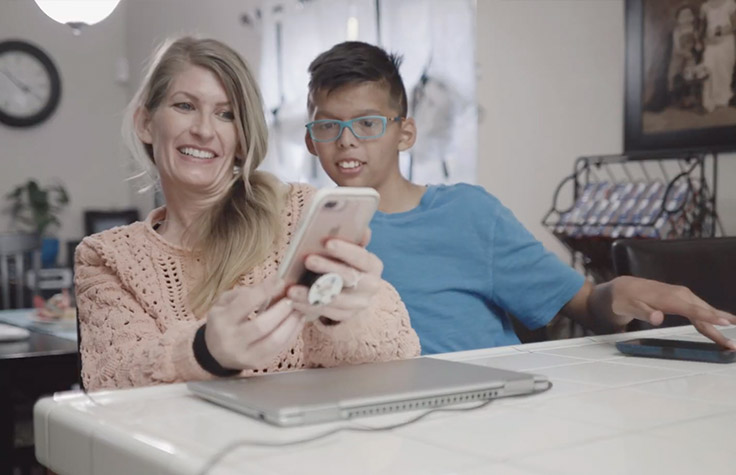

A diagnosis for Donovan
The list of Donovan’s symptoms raised suspicion of more than 20 different conditions and aligned with nearly a dozen medical specialties. After 6 years, whole-genome sequencing identified a variant in the SKI gene, and Donovan was diagnosed with Shprintzen-Goldberg syndrome.

End the odyssey
It can take years to find a diagnosis for patients with a suspected genetic disorder. See how whole-genome sequencing is shortening the diagnostic odyssey and becoming increasingly available.
Visit SiteGenomic Methods for Rare Disease Diagnosis
Whole-Genome sequencing for rare diseases
Whole-genome sequencing is the most comprehensive method for rare disease testing. It examines the entire genome and has the capability to assess variants in both coding and noncoding regions of the genome.12-19
Whole-exome sequencing for rare diseases
Whole-exome sequencing evaluates the exons, the coding regions of the genome, for variants associated with disease.9,10,20
Targeted sequencing for rare diseases
Targeted sequencing analyzes specific genes associated with a rare disease or rare disease family.
Chromosomal microarrays
Chromosomal microarray (CMA) technology identifies large chromosomal variation and specific, well-described variants across the genome.
Compare genomic methods for rare disease diagnosis
Diagnostic yield is the statistic most commonly used to compare genomic testing methods for rare disease. This refers to the likelihood that a test will provide information needed to establish a molecular diagnosis. Diagnostic yield can vary significantly depending on the patient population being studied and the inclusion criteria.
Whole-Genome Sequencing
In most studies, whole-genome sequencing (WGS) shows the highest diagnostic yield of all methods. It broadly covers the genome (> 97%) and is capable of detecting multiple variant types (single nucleotide variants, indels, structural variants, copy number variants, repeat expansions, mitochondrial variants, and paralogs).10-17
Whole-Exome Sequencing
Whole-exome sequencing (WES) has the next highest diagnostic yield. Compared to WGS, WES has less genomic coverage (covering ~1.5% of the genome) and detects fewer variant types. However, WES is less expensive than WGS and generally has higher rates of reimbursement.7–8, 18
Targeted Sequencing
Targeted sequencing for rare disease assesses specific genes. The largest panels cover less than 0.5% of the genome.
Chromosomal Microarrays
Chromosomal microarray methods cover < 0.01% of the genome. CMA focuses specifically on regions of the genome with well-characterized disease-causing variants. CMA tends to have significantly lower diagnostic yield than WES and WGS.7

Maximize your diagnostic potential
Advances in genomic testing are leading to answers faster than ever before. Learn how whole-genome sequencing has been shown to impact clinical management and provide increased actionability.
Download BrochureOnline Course: Clinical Sequencing in Rare Disease
This course offers an overview of pediatric rare disease, available testing options, and clinical implementation of genomic sequencing. It may be relevant to laboratory providers, healthcare providers, healthcare organizations, and others interested in a review of genomics in the rare disease population. This course was made possible through an educational grant from Illumina.
View Course Details
Featured News in Rare Disease Genomics

Making genomics available to all
Making genomics available to all is critical in realizing its potential to save and improve lives. That’s why we are driving down the cost of sequencing, expanding access to advanced technology, and increasing the diversity of genomics data.

Project baby bear results
A pilot program for babies in intensive care showed improved clinical outcomes, a better care experience, and reduced net healthcare expenditures.
Improving care for critically ill infants with whole-genome sequencing
Learn about the NICUSeq study, a randomized time-delayed trial involving five children’s hospitals. This webinar discusses key findings and considerations for implementing diagnostic whole-genome sequencing to serve an acute care setting.
Watch WebinarThe Impact of Precision Genomics on Rare Disease
Escape from limbo land
In this podcast episode, Heather Renton of Syndromes Without A Name (SWAN) Australia discusses her daughter’s rare disease, the diagnostic odyssey, and the impact of NGS.
NICUSeq infographic
Download this infographic to see the impact of whole-genome sequencing on the clinical management of acutely ill newborns with suspected genetic disease.
One mother’s quest for a diagnosis
As a small child, Shubao suffered from hypertonia. Whole-genome sequencing identified a variant in both copies of his PDHX gene, resulting in a tailored treatment. He showed almost immediate improvement.
References
- Nguengang Wakap, S., Lambert, D.M., Olry, A. et al. Estimating cumulative point prevalence of rare diseases: analysis of the Orphanet database. Eur J Hum Genet. 2020;28:165–173. https://doi.org/10.1038/s41431-019-0508-0
- Ferreira CR. The burden of rare diseases. Am J Med Genet A. 2019;179(6):885-892. doi:10.1002/ajmg.a.61124
- Walker CE, Mahede T, Davis G, et al. The collective impact of rare diseases in Western Australia: an estimate using a population-based cohort. Genet Med. 2017;19(5):546-552. doi:10.1038/gim.2016.143
- Global Commission to End the Diagnostic Odyssey for Children with a Rare Disease. 2019.
- Rare Disease Impact Report: Insights from patients and the medical community. globalgenes.org/wp-content/uploads/2013/04/ShireReport-1.pdf.
- Bick D, Jones M, Taylor SL, Taft RJ, Belmont J. Case for genome sequencing in infants and children with rare, undiagnosed or genetic diseases. J Med Genet. 2019;56(12):783-791. doi:10.1136/jmedgenet-2019-106111.
- Clark MM, Stark Z, Farnaes L, et al. Meta-analysis of the diagnostic and clinical utility of genome and exome sequencing and chromosomal microarray in children with suspected genetic diseases. NPJ Genom Med. 2018;3:16. https://doi.org/10.1038/s41525-018-0053-8
- Vissers LE, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet. 2016;17(1):9-18. doi:10.1038/nrg3999
- ACMG Board of Directors. Clinical utility of genetic and genomic services: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2015;17(6):505-507. doi:10.1038/gim.2015.41
- Lionel AC, Costain G, Monfared N, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2018;20(4):435-443. doi:10.1038/gim.2017.119
- Sanghvi RV, Buhay CJ, Powell BC, et al. Characterizing reduced coverage regions through comparison of exome and genome sequencing data across 10 centers. Genet Med. 2018;20(8):855-866. doi:10.1038/gim.2017.192
- Dolzhenko E, van Vugt JJ, Shaw RJ, Bekritsky, et al. Detection of long repeat expansions from PCR-free whole-genome sequence data. Genome Res. 2017;27(11): 1895-1903. doi: 10.1101/gr.225672.117.
- Gross, A.M., Ajay, S.S., Rajan, V. et al. Copy-number variants in clinical genome sequencing: deployment and interpretation for rare and undiagnosed disease. Genet Med. 2019;21:1121–1130. https://doi.org/10.1038/s41436-018-0295-y
- Alfares A, Aloraini T, Subaie LA, et al. Whole-genome sequencing offers additional but limited clinical utility compared with reanalysis of whole-exome sequencing. Genet Med. 2018;20(11):1328-1333. doi:10.1038/gim.2018.41
- Lindstrand A, Eisfeldt J, Pettersson M, et al. From cytogenetics to cytogenomics: whole-genome sequencing as a first-line test comprehensively captures the diverse spectrum of disease-causing genetic variation underlying intellectual disability. Genome Med. 2019;11(1):68. Published 2019 Nov 7. doi:10.1186/s13073-019-0675-1
- Chen X, Sanchis-Juan A, French CE, et al. Spinal muscular atrophy diagnosis and carrier screening from genome sequencing data. Genet Med. 2020;22(5):945-953. doi:10.1038/s41436-020-0754-0
- Chen X, Schulz-Trieglaff O, Shaw R, et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32(8):1220-1222. doi:10.1093/bioinformatics/btv710
- Srivastava S, Love-Nichols JA, Dies KA, et al. Meta-analysis and multidisciplinary consensus statement: exome sequencing is a first-tier clinical diagnostic test for individuals with neurodevelopmental disorders. Genet Med. 2019;21(11):2413-2421. doi:10.1038/s41436-019-0554-6