
NovaSeq X Series ordering
Advanced chemistry, optics, and informatics combine to deliver exceptional sequencing speed and data quality, outstanding throughput, and scalability.
Robust methylation profiling microarray providing extensive coverage of CpG islands, genes, and enhancers. Ideal for genetic and rare disease research, cancer research, and classification.
Sample throughput
Number of samples
Number of markers
The Infinium MethylationEPIC v2.0 kit is a genome-wide methylation analysis tool targeting ~930K unique methylation sites in the most biologically significant regions of the human methylome.
Genome-wide coverage at a relative affordable cost enables scalable discovery of powerful biological insights into disease mechanisms.
Illumina methylation BeadChips have powered > 10 years of groundbreaking research, with most published epigenome-wide association studies (EWAS) to date completed on MethylationEPIC or its predecessor. Harness methylation data in data repositories like the Gene Expression Omnibus (GEO) or The Cancer Genome Atlas (TCGA).
In addition to whole blood, Methylation EPICv2.0 is also compatible with FFPE sample types, enabling studies to be completed on large biorepositories of tumors.
| Assay type | Infinium HD Methylation |
|---|---|
| Automation capability | Automated array loader |
| Description | The Infinium MethylationEPIC v2.0 BeadChip is a genome-wide methylation analysis tool with enhanced, expert-selected functional content. The kit is processed on the iScan or NextSeq 550 Systems and follows a straightforward, user-friendly workflow that does not require sample pooling and indexing. |
| Input quantity | 250 ng DNA |
| Instruments | NextSeq 550 System, NextSeq 550Dx in Research Mode, iScan System |
| Method | Methylation array |
| Nucleic acid type | DNA |
| Number of markers | ~ 930k |
| Number of samples | 8 samples per array |
| Sample throughput | 3024 samples per week on a single iSCAN |
| Specialized sample types | Blood, FFPE tissue |
| Species category | Human |
| Technology | Microarray |
| Variant class | Differentially methylated cytosines |
Requires purchase of a kit that matches the number of samples you run at a time. Please note that each kit is processed as a single batch and is not designed to be divided across multiple experiments. If you are processing several low-throughput batches, order multiple small kits. The kits contain all required reagents for performing methylation screening, except for the bisulfite conversion kits which should be purchased separately from Zymo Research.
Infinium MethylationEPIC v2.0 BeadChip is a robust microarray that enables discovery of methylation-based biomarkers for research applications in cancer, genetic diseases, aging, and molecular epidemiology.
Infinium MethylationEPIC v2.0 Kit
Infinium HD Methylation Assay
Genome-wide methylation profiling studies with microarray and NGS technologies can provide novel insights into the functional impact of methylation.
Complex and genetic disease research
Comprehensive array and next-generation sequencing solutions to accelerate research of various genetic complex diseases.
Explore the benefits of next-generation sequencing and microarrays for analyzing altered methylation patterns and other epigenetic changes in cancer.
| Infinium MethylationEPIC v2.0 Kit | Infinium HTS iSelect Methyl Custom BeadChip | Infinium Methylation Screening Array-48 Kit | Infinium Mouse Methylation BeadChip | |
|---|---|---|---|---|
| Automation capability | Automated array loader | Automated array loader | Automated array loader, Liquid handling robot(s) | Automated array loader |
| Description | The Infinium MethylationEPIC v2.0 BeadChip is a genome-wide methylation analysis tool with enhanced, expert-selected functional content. The kit is processed on the iScan or NextSeq 550 Systems and follows a straightforward, user-friendly workflow that does not require sample pooling and indexing. | Using the DesignStudio Assay Design Tool, this product allows labs to create high-throughput, custom, 24- or 48-sample BeadChips to efficiently and accurately assess methylation for up to 100,000 CpG sites in human and select nonhuman samples. | Highly scalable and affordable methylation analysis of known common disease associations for population health applications. Ideal for studies >1,000 samples and up to millions of samples. | |
| Input quantity | 250 ng DNA | 250 ng DNA | 50 ng DNA | 250 ng DNA |
| Instruments | NextSeq 550 System, NextSeq 550Dx in Research Mode, iScan System | iScan System | iScan System | iScan System |
| Method | Methylation array | Methylation array | Methylation array | Methylation array |
| Nucleic acid type | DNA | DNA | DNA | DNA |
| Number of samples | 8 samples per array | 24 samples per array | 48 samples per array | 12 samples per array |
| Specialized sample types | Blood, FFPE tissue | Blood, Cell-free DNA, Saliva | Low-input samples | Blood |
| Species category | Human | Mouse, Human | Human | Mouse |
| Technology | Microarray | Microarray | Microarray | Microarray |
| Variant class | Differentially methylated cytosines | Differentially methylated cytosines | Differentially methylated cytosines | Differentially methylated cytosines |
Library Prep and Array Kit Selector
Find the right sequencing library preparation kit or microarray for your needs. Filter by method, species, and more. Compare, share, and order kits.
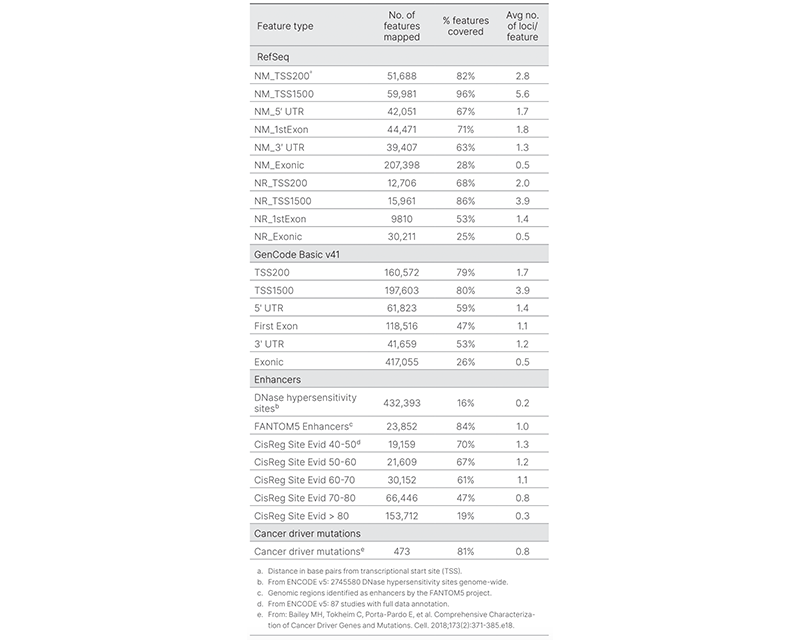

New and enhanced content enables future epigenetic discoveries.
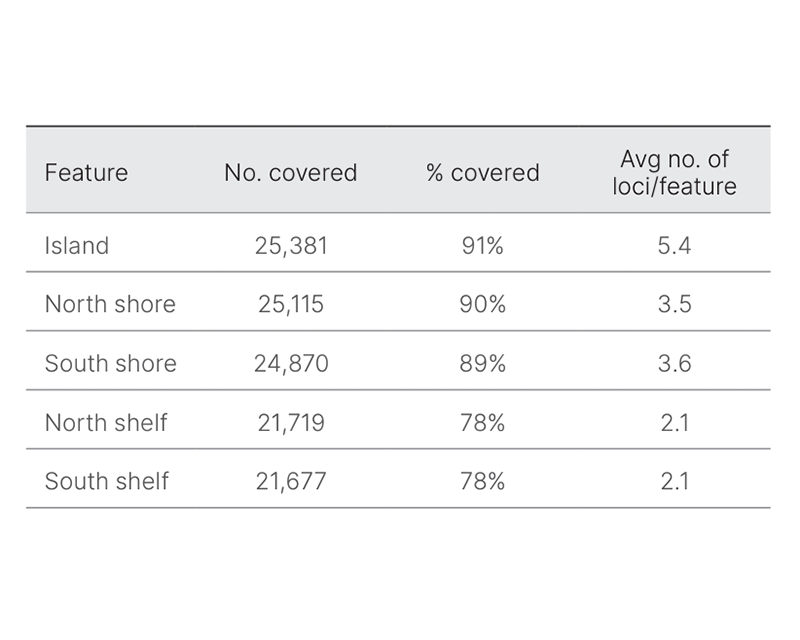

CpG islands and exons that were covered insufficiently on the Infinium MethylationEPIC v1.0 BeadChip are covered thoroughly with additional probes.


FFPE tissue samples show robust performance with a modified version of the Infinium Methylation HD Assay. For precious samples and optimal sample integrity, the Infinium FFPE QC and DNA Restoration Kits are recommended.
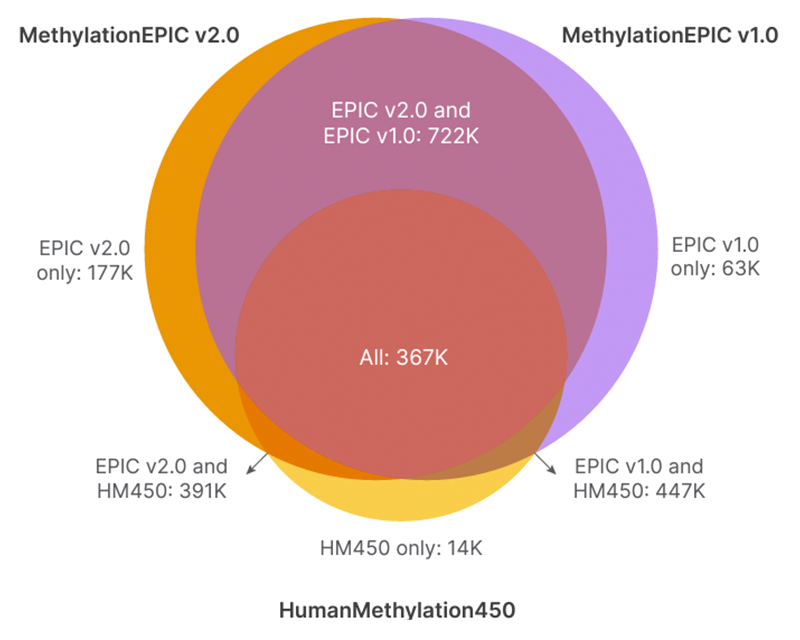

The Infinium MethylationEPIC v2.0 BeadChip builds upon the existing CpG backbones of the Infinium MethylationEPIC v1.0 and HumanMethylation450 BeadChips.
Inf MethylationEPIC V2.0 Kit (8 Spl)
20087706
Includes one, 8-sample BeadChip and reagents for amplifying, fragmenting, hybridizing, labeling, and detecting 8 DNA samples. Requires 250 ng DNA for input. Each kit is processed as a single batch. Purchase bisulfite conversion kits separately.
List Price:
Discounts:
Inf MethylationEPIC V2.0 Kit (16 Spl)
20087707
Includes two, 8-sample BeadChips and reagents for amplifying, fragmenting, hybridizing, labeling, and detecting 16 DNA samples. Requires 250 ng DNA for input. Each kit is processed as a single batch. Purchase bisulfite conversion kits separately.
List Price:
Discounts:
Inf MethylationEPIC V2.0 Kit (32 Spl)
20087708
Includes four, 8-sample BeadChips and reagents for amplifying, fragmenting, hybridizing, labeling, and detecting 32 DNA samples. Requires 250 ng DNA for input. Each kit is processed as a single batch. Purchase bisulfite conversion kits separately.
List Price:
Discounts:
Inf MethylationEPIC V2.0 Kit (96 Spl)
20087709
Includes 12, 8-sample BeadChips and reagents for amplifying, fragmenting, hybridizing, labeling, and detecting 96 DNA samples. Requires 250 ng DNA for input. Each kit is processed as a single batch. Purchase bisulfite conversion kits separately.
List Price:
Discounts:
Inf MethylationEPIC-8+ v2.0 Kit (16 Spl)
20097750
Semi-custom EPIC v2 containing a custom selection of 3,000-10,000 attempted beadtypes on top of EPIC v2. Kit contains 2 beadchips with reagents for processing 16 samples.
Inf MethylationEPIC-8+ v2.0 Kit (32 Spl)
20097751
Semi-custom EPIC v2 containing a custom selection of 3,000-10,000 attempted beadtypes on top of EPIC v2. Kit contains 4 beadchips with reagents for processing 32 samples.
Inf MethylationEPIC-8+ v2.0 Kit (96 Spl)
20097752
Semi-custom EPIC v2 containing a custom selection of 3,000-10,000 attempted beadtypes on top of EPIC v2. Kit contains 12 beadchips with reagents for processing 96 samples.
Infinium® Assay: An Introduction – Customer Site
20015273
Three-day, hands-on instruction at customer site to familiarize users with the essential steps in the Infinium protocol. Course provides hands-on training in sample and BeadChip preparation, sample scanning using the iScan System, and primary data evaluation and analysis using array analysis software.
Showing of
Product
Qty
Unit price
Product
Catalog ID
Quantity
Unit price
The Infinium Methylation EPIC v2.0 BeadChip Kit leverages powerful Infinium chemistry to detect cytosine methylation at CpG islands based on highly multiplexed genotyping of bisulfite-converted genomic DNA. Learn more about quantitative array-based methylation measurement.
This kit uses the Illumina Methylation Assay, a microarray technology that provides quantitative methylation measurement at the single-CpG-site level, powering research that examines the biological role of DNA methylation in normal and diseased cells. Learn more about the Illumina Methylation Assay.
The Infinium MethylationEPIC v2.0 BeadChip Kit contains 186K new probes that target known enhancers, super-enhancers, CTCF-binding sites, and open regions of chromatin associated with primary tumors identified by ATAC-Seq and ChIP-seq experiments. Learn about additional improvements in the Infinium MethylationEPIC v2.0 data sheet.
Illumina offers user friendly tools to analyze Infinium controls built into the MethylationEPIC BeadChip. For more information visit the Methylation Array Data Analysis Tips Page.
The Zymo EZ DNA Methylation Lightning Kit is the recommended bisulfite conversion kit for the Infinium workflow. Third party bisulfite conversion kits must be purchased separately. Learn more about validated bisulfite conversion kits for Infinium Methylation BeadChips.
The scan takes ~18–37 minutes per BeadChip.
Yes, the Infinium MethylationEPIC v1.0 and HumanMethylation450 BeadChip kits have been discontinued. The Infinium MethylationEPIC v2.0 Kit, built upon the same reliable Infinium chemistry foundation, offers enhanced, expert-selected content and is the recommended replacement for both kits.
Reach out for information about our products and services, or get answers to questions about our technology.
Your email address is never shared with third parties.