FMRI achieves higher efficiency in genomic research with Illumina’s new exome enrichment.
Introduction
Ms. Sookyoung Kim is the Chief Researcher of Future Medicine Research Institute (FMRI) located in CHA University Bundang Hospital (CUBH) in Gyeonggi-do, South Korea, a major branch hospital of CHA University Medical Center. FMRI was recently established to support researchers across CHA University Medical Center. FMRI has plans to provide various NGS service using NextSeq™ 550Dx in RUO mode. In particular, we asked Ms. Kim about her recent experience using Illumina DNA Prep with Exome 2.0 Plus Enrichment.
CHA University Medical Center is comprised of 86-branch hospitals and institutes which has specialties in women's healthcare and reproductivity in 7 countries. In 1986, it became the first in Korea to succeed the in-vitro fertilization (IVF) procedure and now it is one of the most reputable hospitals for IVF in Korea. In recent years, it has been striving to develop cures for intractable disease and new drugs as a top-notch, research focused medical institution.
Illumina DNA Prep with Exome 2.0 Plus Enrichment
Illumina DNA Prep with Exome 2.0 Plus Enrichment is a component of Illumina’s comprehensive exome solution, starting from library preparation, high-performance sequencing instruments and award-winning secondary data analysis using DRAGEN™ Enrichment App.
Illumina DNA Prep with Exome 2.0 Plus Enrichment features shorter hands-on preparation time, improved target region coverage and a cost-effective panel size. Not only that, but it also enables streamlined implementation of a single platform from a single vendor – from library preparation to data analysis.
Q: Was the result with Illumina DNA Prep with Exome 2.0 Plus Enrichment satisfactory to implement Exome library prep at FMRI?
Answer: Overall, the result was satisfactory. We saw high target coverage, best uniformity (over 98%), good on-target rate (over 93%), and low duplication rate (around 10%) increasing the production efficiency of sequencing data as well as reduce the sequencing running cost.
Q: Do you think Illumina DNA Prep with Exome 2.0 Plus Enrichment can consolidate current gene testing by non-NGS method and/or small gene panel testing?
Answer: Since Illumina DNA Prep with Exome 2.0 Plus Enrichment has significant advantages in terms of accuracy and speed of analysis, it is a worthy replacement for the current gene testing. When testing multiple targets, it is especially advantageous compared to the non-NGS method or small gene panel testing. However, somatic mutation analysis requires higher read depth and exome for somatic mutation is considered suitable only for new biomarker discovery purposes because of sequencing costs. On the other hand, it is important for the researchers in the laboratory to access equipment that is easy to operate or workflow that is easy to follow. Compared to the existing library preparation protocol, the workflow of Illumina DNA Prep with Exome 2.0 Plus Enrichment was simple and data reproducibility was also great.
Since Illumina DNA Prep with Exome 2.0 Plus Enrichment has significant advantages in terms of accuracy and speed of analysis, it is a worthy replacement for the current gene testing.
Q: What was the advantage of using DRAGEN™ compared to the open-source 2nd analysis currently use?
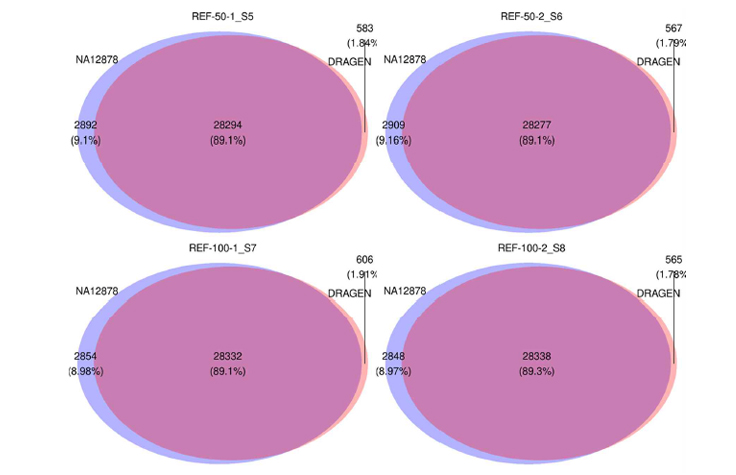
Answer: It is impressive that the execution time of the NGS pipeline was very much reduced using DRAGEN. The BaseSpace Sequence Hub (BSSH) command line interface (CLI) also significantly reduces the time to upload files to the cloud, which appears to be a huge advantage. Also, a user-friendly interface provides convenience for non-experts to use this platform. In terms of our analysis result, the target coverage at all depths, mean target coverage depth, and percent Q30 of the DRAGEN are higher than that of our analysis pipeline in all samples. When comparing the variants of the DRAGEN and our analysis pipeline, approximately 90% of SNPs were consistent in all samples, and approximately 65% of INDEL variants were consistent. We can regard sufficient coverage depth and consistent callings level (Fig.1).
Q: What were the best aspects and the biggest opportunities for your overall workflow, including library prep with Illumina DNA Prep with Exome 2.0 Plus Enrichment, and sequencing with the NextSeq™ 550Dx system in RUO mode?
Answer: The library preparation process has been simplified. The hands-on time has been reduced and the quality of the result is satisfactory. However, since the purity of DNA is important when using the enzymatic tagmentation method, many samples do not pass this standard, so it seems necessary to provide additional guidelines (e.g. increase input amount and pre-PCR cycle, adjust pooling ratio, and etc.).
The library preparation process has been simplified. The hands-on time has been reduced and the quality of the result is satisfactory.
Q: How do you feel about the support provided by Illumina throughout the process of this evaluation?
Answer: We appreciated the detailed technical advice on how to use BSSH, and friendly resolution of various inquires Illumina provided. Members of Illumina Korea I met had an affection for Illumina products and wanted to accurately convey the strengths of the products. Overall, the support was quite impressive. I hope that Illumina continues to support various genomic research with its products of excellent performance, and continue to make advancements in these technologies. I believe our success at FMRI is interdependent with Illumina’s progress.
I believe our success at FMRI is interdependent with Illumina’s progress.
Learn more about the products mentioned in this case study:
DNA Prep with Exome 2.0 Plus Enrichment